RSE Stories: An Interview With Dr Juan Nunez Iglesias
Posted by Paula Martinez on 1 July 2022
This post was originally published as a news item on the Australian Research Data Commons website (1 July, 2022).
As part of our Research Software Agenda for Australia, the ARDC is working with the research community to shape better research software in order to recognise it as a first-class output of research.
This interview is part of a series about research software engineers. Each month we talk with a research software engineer about their experiences and best-practice tips in creating, improving and sustaining software for research.
Continuing the series, we talked to Dr Juan Nunez-Iglesias, a Senior Research Fellow and Group Leader at the Monash Biomedicine Discovery Institute, at Monash University. Find out about his journey into the world of open source software and the amazing collaborators he met on his way.
What do you do in your role as a research software engineer?
I develop methods and tools in image analysis and computer vision across scientific domains (including bioimaging, remote sensing/satellite imagery, and materials science), particularly for 3D and higher-dimensional images, which are common in microscopy and other bioimaging techniques.
I work almost exclusively in Python. When I need higher performance I reach for Numba, a compiler built for scientific computing that makes Python code run faster, with little effort.
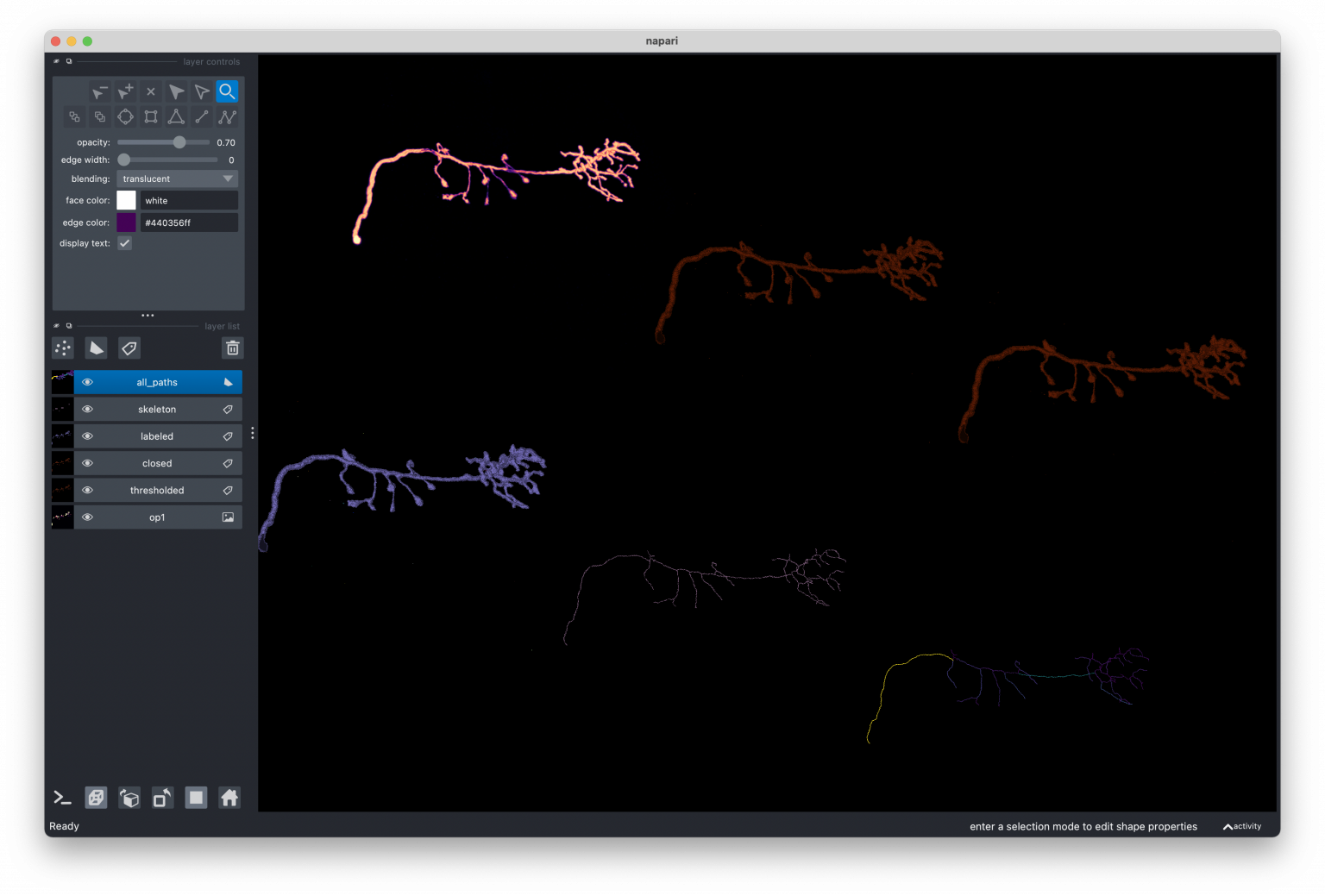
My main projects are napari, an n-dimensional image viewer for Python that I co-founded with Loïc Royer (Chan Zuckerberg Biohub), and scikit-image, an image analysis and processing library started by my friend and mentor Stéfan van der Walt (UC Berkeley, formerly Stellenbosch University). Both are research software projects that accelerate biomedical research and serve the larger scientific community worldwide. For example, scikit-image is being used in diverse research domains - for example to detect lung cancer, measure deforestation, and in archaeology to transcribe clay tablets from the 7th century BC. How cool is that!
How did you come to work as a research software engineer (RSE)?
I think I am like most RSEs in that my path was very roundabout! I certainly did not set out to be an RSE, though the RSE-eeds were planted early.
For my undergrad, I enrolled in the biomedical science degree at Melbourne University. I wanted to study ageing, but I also really liked computers so I enrolled in a bunch of subjects from the computer science department, and I really enjoyed them! During my PhD I shifted further on the computational spectrum after I realised I hated lab work. I was lucky because I happened to go to the University of Southern California, which was home to the first PhD program in computational biology in the world, founded in 1982 by Michael Waterman. So my transition from the bench to coding at the desktop was extremely well supported.
Back then I was working on gene expression measurements (using microarrays), and I used lots of R and some C++. During my PhD I discovered version control with subversion, which was my first inkling that there was more to writing good research software than writing it.
I then did a postdoc at Janelia Research Campus working on image segmentation. This was a software engineering-intensive project and it took me to the SciPy 2012 conference, which is where I really got into open source. It was at SciPy that I realised that there were scientists, like you and me, who go that extra mile to not reinvent the wheel, and to make their library maximally re-usable. They also used standard workflows using GitHub to work collaboratively. That appealed to me a lot!
After I moved back to Australia, I joined the Life Sciences Computation Centre (which later became Melbourne Bioinformatics), where I was the only person working on imaging data and image analysis. In late 2018, while working at Monash Micro Imaging, the core light microscopy facility at Monash, I was fortunate to be granted a 3-year fellowship (since extended to 5 years) from the Chan Zuckerberg Initiative (CZI) to support some open source maintenance work that was going into scientific image analysis software — Fiji, CellProfiler, and scikit-image. The fellowship enabled me to move to a faculty role at the Monash Biomedicine Discovery Institute.
Now, with the fellowship, I do have a lot of independence to focus on things that traditionally have not been valued in academia, such as writing quality software, maintaining it and helping others maintain it and adapt it for their needs, and therefore growing the community around that software.
I think that universities are missing a trick in not investing more in this type of work, which would give advantages to the universities through better, tailor-made software to support their research; better, more transferable skills and training for their students and employees; and increased reputation.
What best practices do you implement for quality software?
The number one thing is: never code alone.
This is hard, but you should really make the effort to make it happen. For example, find a buddy in your university who works on a different problem, in a different lab, then work together half time on one project and half time on the other. It will seem, at first, that you are slowing down to half speed. But, by preventing errors and designing software that at least 2 people can use, you will in the end save time. Co-developing helps you avoid having to rewrite your software multiple times because your initial design will almost always be flawed — no matter how experienced you are.
Tell us about a project you have enjoyed working on
I really liked working on skan (skeleton analysis), because it was a smaller project with a very specific purpose. I worked in close collaboration with the biophysicist who was the primary user, which allowed me to experiment with technologies such as Numba, and push the performance as far as I could. I gave a talk about this at SciPy 2019, where I demonstrated how I was hitting the main memory bandwidth of my computer and the community response helped me improve it.
Bigger projects such as scikit-image and
napari are also interesting, but in a different
way. Because more people rely on them, stability and reliability matter
a lot more than being on the cutting edge, which gives you less freedom
to experiment. Also, serving a bigger community means more time spent
reviewing others' requests and contributions, rather than coding. So
skan feels a bit like a playground in comparison!
Who do you work with?
I have 2 PhD students who are amazing. Abigail McGovern, who works on bioimage informatics, and Draga Doncila Pop, who works on human-in-the-loop semi-automatic 3D tracking. In addition to her PhD, Draga works as a consultant for CZI on napari development.
Further afield, my work as an open source maintainer gives me the opportunity to collaborate with amazing folks around the world, including Talley Lambert at Harvard Medical School, Andy Sweet at CZI, and Kevin Yamauchi at ETH Zurich, among many others. Napari now has 118 direct contributors, so there's far too many interesting folks I work with that I don't have space to mention!
Which research software communities do you recommend?
For all your bioimage analysis needs, Pythonic or otherwise, go to forum.image.sc. This forum represents more than 60 open source software packages, including Fiji, CellProfiler, scikit-image, and napari. It's full of extremely helpful people, including the authors of all those programs and libraries, and one of the best places to learn both image analysis principles, and how to use specific bits of software, straight from the horse's mouth, so to speak.
Napari has its own Zulip chat room. Zulip is a free and open source alternative to Slack with much better support for asynchronous communication. The napari Zulip is fantastic, because napari now has core developers scattered all around the globe, so you can usually find someone to chat with about your napari issues 24/7.
You can connect with Juan via Twitter and GitHub or Gitlab.
Stay tuned for our next interview in the Shaping Research Software series, coming out in September.
If you’d like to be part of the growing community of research software engineers in Australia, become a member of the RSE Association of Australia and New Zealand (RSE-AUNZ) (it’s free!).
Learn more about the ARDC’s Research Software Agenda for Australia.
The ARDC is funded through the National Collaborative Research Infrastructure Strategy (NCRIS) to support national digital research infrastructure for Australian researchers.
Dr Juan Nunez-Iglesias was interviewed by Dr Paula Andrea Martinez, ARDC. Edited by Jo Savill (ARDC).
